Technology
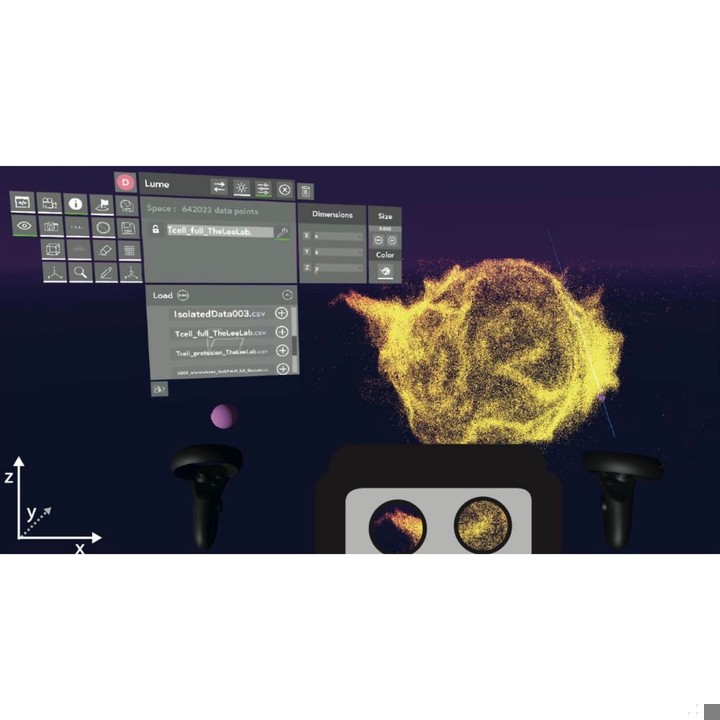
vLUME (pronounced "volume") is a virtual reality software package designed to visualize and analyze large three-dimensional single-molecule localization microscopy (3D-SMLM) datasets. SMLM techniques like photoactivated localization microscopy (PALM) and stochastic optical reconstruction microscopy (STORM) can image biological structures at nanometer resolution by determining the locations of individual fluorescent molecules. However, the analysis of the resulting datasets consisting of millions of localization points poses challenges. vLUME leverages virtual reality technology to allow intuitive visualization and quantification of complex 3D SMLM data.
The vLUME software loads existing 3D-SMLM datasets where each fluorescent molecule is defined by its x, y, and z coordinates. These datasets can come from various SMLM techniques and be processed by different localization algorithms. Once loaded into the virtual environment, the millions of molecule positions appear as points reconstructing the original structure in three dimensions. Users wearing a head-mounted VR display and controllers can then visually explore the nanoscale biological architecture from any angle by walking around and manipulating the visualization.
Key features of vLUME include:
1) Intuitive visualization and navigation of datasets consisting of up to millions of localizations in an immersive 3D space. This facilitates identifying artifacts in the data and comparing datasets processed with different algorithms.
2) Segmentation of regions of interest by using VR controllers to draw 3D selection boxes around subsets of points representing particular structural elements. These selections can be isolated, visualized separately, and exported for further analysis. For example, an individual microtubule filament can be extracted from a complex microtubule network.
3) Custom analysis of selected regions of interest by running user-defined scripts. Quantifications like cluster analysis and nearest neighbor distances can provide spatial statistics revealing organizational principles of nanoscale biological assemblies. Results are displayed directly within the VR environment.
4) Creation of fly-through videos along user-defined paths, allowing effective communication of scientific discoveries.
Under the hood, vLUME is built on the Unity 3D gaming engine, providing high-performance interactive rendering of large point clouds. It runs on consumer VR hardware like the Oculus Rift and HTC Vive. The software interface is tailored to life scientists without need for VR or programming expertise.
vLUME aims to make 3D-SMLM data more accessible and quantifiable compared to analysis on flat screens with standard visualization software. The immersive visualization can provide intuition about nanoscale topology that is difficult to convey in two dimensions. Simple tools for segmentation and scripting also enable quantitative characterization of complex biological structures. By facilitating new ways of seeing and understanding 3D-SMLM datasets, vLUME has the potential to drive new breakthroughs and applications of super-resolution microscopy.
Publications featuring vLume

|
vLUME - 3D virtual reality for single-molecule localization microscopy Alexander Spark, Alexandre Kitching, Daniel Esteban-Ferrer, Anoushka Handa, Alexander R. Carr, Lisa-Maria Needham, Aleks Ponjavic, Ana Mafalda Santos, James McColl, Christophe Leterrier, Simon J. Davis, Ricardo Henriques, Steven F. Lee Paper published in Nature Methods, October 2020 Technologies: Nuclear-Pores as references, QuickPALM and vLume Funded by: BBSRC and Wellcome Trust News: Technology Networks, La Razon (ESP), ZAP and Medium US Blogs: Qubit, , Tech Explorist and Whats New DOI: 10.1038/s41592-020-0962-1 |
|
Funding contributing to vLume

|
VirusAwareScopes - Machine Learning-Driven Adaptive Microscopy for Long-Term Viral Infection Studies Ricardo Henriques Alias: VirusAwareScopes Funded by: La Caixa Foundation - Health Research Duration: November 2025 - October 2028 |

|
VP-CLEM-KIT - a pipeline for democratising volumetric visual proteomics Lucy Collinson, Ricardo Henriques, Paul French Funded by: CZI - Visual Proteomics Imaging Duration: December 2021 - June 2024 Publications: 22 |

|
Enabling Live-Cell 4D Super-Resolution Microscopy Guided by Artificial Intelligence Ricardo Henriques Alias: SelfDriving4DSR Funded by: ERC - Consolidator Duration: July 2021 - June 2026 Publications: 38 |

|
Unveiling live-cell viral replication at the nanoscale Ricardo Henriques Funded by: EMBO - Installation Grant Duration: January 2021 - December 2025 Publications: 34 |