Technology
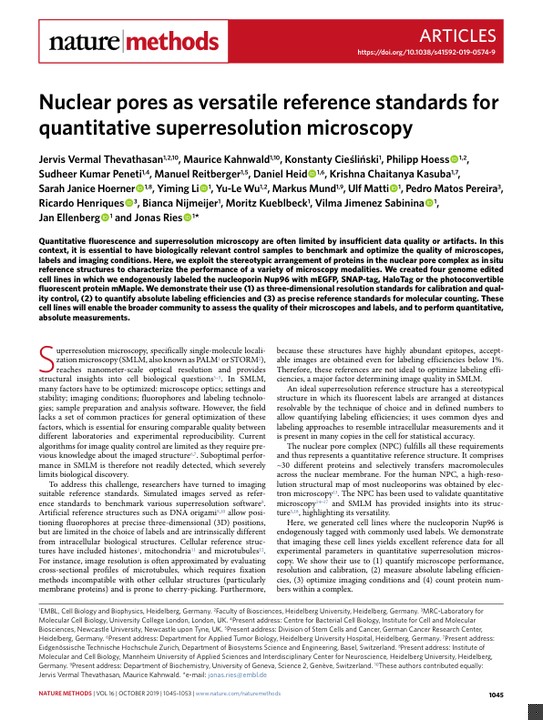

The nuclear pore complex (NPC) is a large protein complex that forms channels through the nuclear envelope, allowing selective transport of molecules between the nucleus and cytoplasm. Recent advances in super-resolution microscopy have enabled imaging the NPC structure with nanometer precision.
The NPC has a stereotypical structure with well-defined dimensions, making it an ideal candidate to serve as a reference standard for quantitative microscopy. Specifically, the nucleoporin Nup96 is present in 32 copies per NPC, forming cytoplasmic and nucleoplasmic rings that contain 16 copies each. The corners of these rings contain pairs of Nup96 proteins that are just 12 nm apart, within the resolution range of most super-resolution techniques.
By genetically tagging Nup96 with fluorescent proteins or tags like GFP, HaloTag or SNAP-tag, the distribution of labels can serve as a benchmark to characterize microscope performance. As the architecture of NPCs is conserved across cells, imaging parameters like resolution, calibration and labeling efficiency can be quantified by analyzing the Nup96 label distribution in the rings.
For example, the diameter of the ring structure reports on the resolution. If the corners are not well resolved, it indicates lower resolution compared to a microscope that clearly resolves all eight corners. Similarly, deviations in measured ring diameters or axial distances would reveal inaccuracies in microscope calibration.
As the number of Nup96 copies per NPC is known, the distribution of labels also enables quantifying labeling efficiency both across imaging conditions and between different labels like antibodies or dyes. A lower fraction of visible corners indicates reduced labeling efficiency. Optimizing imaging buffers or antibody selections to maximize visible corners improves effective labeling for that target.
The abundance and consistent geometry of NPCs further aids statistical quantification. As hundreds of nuclear pores are simultaneously imaged at the bottom of the nucleus, variability between cells and experiments is minimized. The flat geometry also simplifies image analysis.
Genetically labeled Nup96 proteins allow the nuclear pore complex to serve as an in situ reference standard. The stereotypical NPC structure enables benchmarking microscope resolution, calibration and labeling efficiencies. As NPCs robustly incorporate different tags like GFP or HaloTag, the same cell line permits optimization across different imaging modes. The availability of multiple labeled copies per cell reduces sampling errors and simplifies analysis. Together, these properties make endogenously tagged Nup96 a versatile tool to enhance precision and reproducibility in quantitative microscopy assays.
Publications featuring Nuclear-Pores as references

|
Expansion and fluctuations-enhanced microscopy for nanoscale molecular profiling of cells and tissues Dominik Kylies, Hannah S. Heil, Arturo G. Vesga, Mario Del Rosario, Maria Schwerk, Malte Kuehl, Milagros N. Wong, Victor G. Puelles, Ricardo Henriques Paper published in Nature Protocols, July 2025 Technologies: NanoJ (), NanoJ-eSRRF (), NanoJ-SQUIRREL (), NanoJ-SRRF (), NanoPyx () and Nuclear-Pores as references Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1038/s41596-025-01178-0 |
|
|
Structural Repetition Detector for multi-scale quantitative mapping of molecular complexes through microscopy Afonso Mendes, Bruno M. Saraiva, Guillaume Jacquemet, João I. Mamede, Christophe Leterrier, Ricardo Henriques Paper published in Nature Communications, July 2025 Technologies: Nuclear-Pores as references and SReD () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, H2021 and H2022 DOI: 10.1038/s41467-025-60709-1 |
|

|
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation Romain F. Laine, Hannah S. Heil, Simao Coelho, Jonathon Nixon-Abell, Angélique Jimenez, Theresa Wiesner, Damián Martínez, Tommaso Galgani, Louise Régnier, Aki Stubb, Gautier Follain, Samantha Webster, Jesse Goyette, Aurelien Dauphin, Audrey Salles, Siân Culley, Guillaume Jacquemet, Bassam Hajj, Christophe Leterrier, Ricardo Henriques Paper published in Nature Methods, November 2023 Technologies: CARE (), NanoJ (), NanoJ-eSRRF (), NanoJ-SQUIRREL (), NanoJ-SRRF () and Nuclear-Pores as references Funded by: CZI, EMBO, ERC, FCT, H2021, H2022, InnOValley and Wellcome Trust News: Photonics.com, The Science Times, Optics.org and Phys.org Blogs: Springer Nature Protocols and Methods Community DOI: 10.1038/s41592-023-02057-w |
|

|
Mapping molecular complexes with super-resolution microscopy and single-particle analysis Afonso Mendes, Hannah S. Heil, Simao Coelho, Christophe Leterrier, Ricardo Henriques Review published in Open Biology, July 2022 Technologies: NanoJ-VirusMapper, Nuclear-Pores as references and ZeroCostDL4Mic () Funded by: EMBO, ERC and Wellcome Trust DOI: 10.1098/rsob.220079 |
|

|
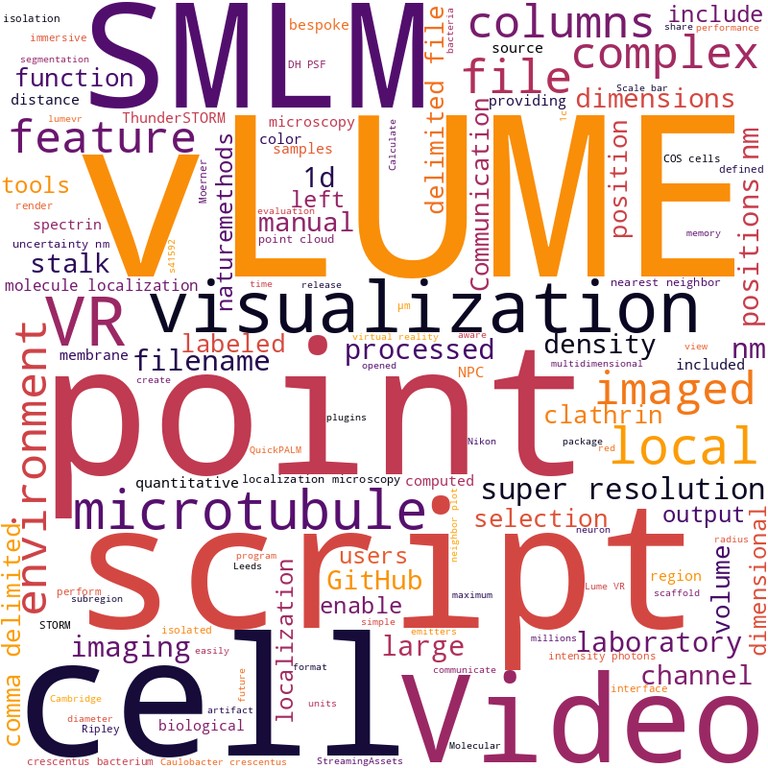
vLUME - 3D virtual reality for single-molecule localization microscopy Alexander Spark, Alexandre Kitching, Daniel Esteban-Ferrer, Anoushka Handa, Alexander R. Carr, Lisa-Maria Needham, Aleks Ponjavic, Ana Mafalda Santos, James McColl, Christophe Leterrier, Simon J. Davis, Ricardo Henriques, Steven F. Lee Paper published in Nature Methods, October 2020 Technologies: Nuclear-Pores as references, QuickPALM and vLume Funded by: BBSRC and Wellcome Trust News: Technology Networks, La Razon (ESP), ZAP and Medium US Blogs: Qubit, , Tech Explorist and Whats New DOI: 10.1038/s41592-020-0962-1 |
|
|
Closed mitosis requires local disassembly of the nuclear envelope Gautam Dey, Siân Culley, Scott Curran, Uwe Schmidt, Ricardo Henriques, Wanda Kukulski, Buzz Baum Paper published in Nature, August 2020 Technologies: CARE (), NanoJ (), NanoJ-SRRF () and Nuclear-Pores as references Funded by: BBSRC and Wellcome Trust News: Nature Asia DOI: 10.1038/s41586-020-2648-3 |
|

|
The cell biologist's guide to super-resolution microscopy Guillaume Jacquemet, Alexandre F. Carisey, Hellyeh Hamidi, Ricardo Henriques, Christophe Leterrier Review published in Journal of Cell Science, June 2020 Technologies: CARE (), NanoJ (), NanoJ-Fluidics (), NanoJ-SRRF () and Nuclear-Pores as references Funded by: BBSRC and Wellcome Trust News: ScienMag and EurekAlert! DOI: 10.1242/jcs.240713 |
|

|
Nuclear pores as versatile reference standards for quantitative superresolution microscopy Jervis Vermal Thevathasan, Maurice Kahnwald, Konstanty Cieśliński, Philipp Hoess, Sudheer Kumar Peneti, Manuel Reitberger, Daniel Heid, Krishna Chaitanya Kasuba, Sarah Janice Hoerner, Yiming Li, Yu-Le Wu, Markus Mund, Ulf Matti, Pedro Matos Pereira, Ricardo Henriques, Bianca Nijmeijer, Moritz Kueblbeck, Vilma Jimenez Sabinina, Jan Ellenberg, Jonas Ries Paper published in Nature Methods, September 2019 Technologies: CARE (), NanoJ (), NanoJ-SQUIRREL (), NanoJ-SRRF () and Nuclear-Pores as references Funded by: BBSRC and Wellcome Trust News: Mirage News DOI: 10.1038/s41592-019-0574-9 |
|
Funding contributing to Nuclear-Pores as references

|
VirusAwareScopes - Machine Learning-Driven Adaptive Microscopy for Long-Term Viral Infection Studies Ricardo Henriques Alias: VirusAwareScopes Funded by: La Caixa Foundation - Health Research Duration: November 2025 - October 2028 |

|
Mapping the early stages of HIV-1 infection by live-cell 4D Super-Resolution Microscopy Hannah Heil Funded by: EMBO - Postdoctoral Fellowships Duration: September 2021 - August 2023 Publications: 5 |

|
Mapping HIV-1 infection by 4D Super-Resolution Microscopy Hannah Heil Funded by: FCT - CEEC Individual Duration: July 2021 - December 2024 Publications: 5 |