Real-Time high-content Super-Resolution Imaging of ES Cell States

Alias: RT-SuperES
Agency: H2022
Type: EIC Pathfinder Open
Principal Investigator: Eran Meshorer
Investigators: Eran Meshorer, Ricardo Henriques, Anna Kreshuk, Sandrine Lévêque-Fort, Nicolas Bourg, Genevieve Almouzni
Start-date: July 2023
End-date: June 2027
Grant Code: 101099654
DOI: 10.3030/101099654
Agency: H2022
Type: EIC Pathfinder Open
Principal Investigator: Eran Meshorer
Investigators: Eran Meshorer, Ricardo Henriques, Anna Kreshuk, Sandrine Lévêque-Fort, Nicolas Bourg, Genevieve Almouzni
Start-date: July 2023
End-date: June 2027
Grant Code: 101099654
DOI: 10.3030/101099654
RT-SuperES is a project that creates a novel microscope equipped with machine learning-based automated decision making, allowing high-content imaging of embryonic stem cells in real-time. By combining conventional and super-resolution imaging, it explores cellular processes during differentiation at a nanoscale level. It will use cutting-edge technologies such as SNAP-tagging, SRRF, SMLM, SIM, NanoJ-Fluidics, and AI algorithms. The project brings together a multidisciplinary team across four countries to establish this groundbreaking and affordable imaging technology.
Technology explored
Supported publications

|
EZInput - A Cross-Environment Python Library for Easy UI Generation in Scientific Computing Bruno M Saraiva, Iván Hidalgo-Cenalmor, António D Brito, Damián Martínez, Tayla Shakespeare, Guillaume Jacquemet, Ricardo Henriques Preprint published in arXiv, January 2026 Technologies: DL4MicEverywhere (), mAIcrobe (), NanoJ-eSRRF (), NanoPyx () and ZeroCostDL4Mic () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, H2021 and H2022 DOI: 10.48550/arXiv.2601.08859 |
|

|
PhotoFiTT - a quantitative framework for assessing phototoxicity in live-cell microscopy experiments Mario Del Rosario, Estibaliz Gómez-de-Mariscal, Leonor Morgado, Raquel Portela, Guillaume Jacquemet, Pedro M. Pereira, Ricardo Henriques Paper published in Nature Communications, December 2025 Technologies: mAIcrobe (), PhotoFiTT () and ZeroCostDL4Mic () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1038/s41467-025-66209-6 |
|

|
SMOLM-LFM - ratiometric single molecule orientation without polarizers Arturo G. Vesga, Luis Aleman-Castaneda, Hannah S. Heil, Sam Daly, Ezra Bruggeman, Steven F. Lee, Sophie Brasselet, Ricardo Henriques Preprint published in bioRxiv, December 2025 Funded by: EMBO, ERC, FCT and H2022 DOI: 10.1101/2025.11.26.690201 |
|

|
mAIcrobe - an open-source framework for high-throughput bacterial image analysis António D. Brito, Dominik Alwardt, Beatriz de P. Mariz, Sérgio R. Filipe, Mariana G Pinho, Bruno M. Saraiva, Ricardo Henriques Preprint published in bioRxiv, October 2025 Technologies: DeepBacs (), mAIcrobe (), Rescale4DL () and ZeroCostDL4Mic () Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1101/2025.10.21.683709 |
|

|
Nanoscale spatial-omics via contrastive embedding of single-molecule localisation data S. Shirgill, L.G. Jensen, D.J. Nieves, M.C. Wales, A. Kaminer, K. Savoye, R. Peters, M. Heilemann, R. Henriques, S.F. Lee, P. Rubin-Delanchy, D.M. Owen Preprint published in bioRxiv, October 2025 Technologies: nano-org Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1101/2025.10.07.679170 |
|

|
CLEM-Reg - an automated point cloud-based registration algorithm for volume correlative light and electron microscopy Daniel Krentzel, Matouš Elphick, Marie-Charlotte Domart, Christopher J Peddie, Romain F Laine, Cameron Shand, Ricardo Henriques, Lucy M Collinson, Martin L Jones Paper published in Nature Methods, September 2025 Technologies: BioImage Model Zoo (), CARE (), DL4MicEverywhere (), Rescale4DL () and ZeroCostDL4Mic () Funded by: CZI, ERC, H2021 and H2022 DOI: 10.1038/s41592-025-02794-0 |
|

|
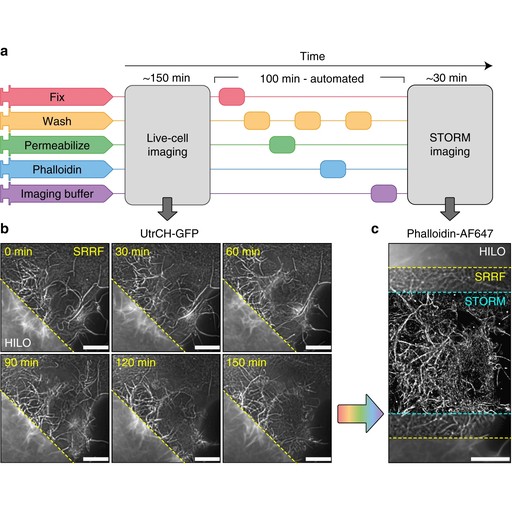
Expansion and fluctuations-enhanced microscopy for nanoscale molecular profiling of cells and tissues Dominik Kylies, Hannah S. Heil, Arturo G. Vesga, Mario Del Rosario, Maria Schwerk, Malte Kuehl, Milagros N. Wong, Victor G. Puelles, Ricardo Henriques Paper published in Nature Protocols, July 2025 Technologies: NanoJ (), NanoJ-eSRRF (), NanoJ-SQUIRREL (), NanoJ-SRRF (), NanoPyx () and Nuclear-Pores as references Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1038/s41596-025-01178-0 |
|
|
Structural Repetition Detector for multi-scale quantitative mapping of molecular complexes through microscopy Afonso Mendes, Bruno M. Saraiva, Guillaume Jacquemet, João I. Mamede, Christophe Leterrier, Ricardo Henriques Paper published in Nature Communications, July 2025 Technologies: Nuclear-Pores as references and SReD () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, H2021 and H2022 DOI: 10.1038/s41467-025-60709-1 |
|
|
Rxiv-Maker - An Automated Template Engine for Streamlined Scientific Publications Bruno M Saraiva, Guillaume Jaquemet, Ricardo Henriques Preprint published in arXiv, June 2025 Technologies: DL4MicEverywhere () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, H2021 and H2022 DOI: 10.48550/arXiv.2508.00836 |
|

|
Nanoscale imaging of biological systems via expansion and super-resolution microscopy Daria Aristova, Dominik Kylies, Mario Del Rosario, Hannah S. Heil, Maria Schwerk, Malte Kuehl, Milagros N. Wong, Ricardo Henriques, Victor G. Puelles Paper published in Applied Physics Reviews, April 2025 Technologies: NanoJ-eSRRF () and NanoJ-SRRF () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1063/5.0240464 |
|

|
The nucleoid of rapidly growing Escherichia coli localizes close to the inner membrane and is organized by transcription, translation, and cell geometry Christoph Spahn, Stuart Middlemiss, Estibaliz Gómez-de-Mariscal, Ricardo Henriques, Helge B. Bode, Séamus Holden, Mike Heilemann Paper published in Nature Communications, April 2025 Technologies: CARE (), DeepBacs () and ZeroCostDL4Mic () Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1038/s41467-025-58723-4 |
|

|
ReScale4DL - Balancing Pixel and Contextual Information for Enhanced Bioimage Segmentation Mariana G. Ferreira, Bruno M. Saraiva, António D. Brito, Mariana G. Pinho, Ricardo Henriques, Estibaliz Gómez-de-Mariscal Preprint published in bioRxiv, April 2025 Technologies: BioImage Model Zoo (), DeepBacs (), DL4MicEverywhere (), NanoPyx (), Rescale4DL () and ZeroCostDL4Mic () Funded by: CZI, EMBO, ERC, H2021 and H2022 DOI: 10.1101/2025.04.09.647871 |
|

|
Efficiently accelerated bioimage analysis with NanoPyx, a Liquid Engine-powered Python framework Bruno M. Saraiva, Inês Cunha, António D. Brito, Gautier Follain, Raquel Portela, Robert Haase, Pedro M. Pereira, Guillaume Jacquemet, Ricardo Henriques Paper published in Nature Methods, January 2025 Technologies: Fast4DReg (), NanoJ (), NanoJ-eSRRF (), NanoJ-SQUIRREL () and NanoPyx () Funded by: CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1038/s41592-024-02562-6 |
|

|
This Microtubule Does Not Exist - Super‐Resolution Microscopy Image Generation by a Diffusion Model Alon Saguy, Tav Nahimov, Maia Lehrman, Estibaliz Gómez‐de‐Mariscal, Iván Hidalgo‐Cenalmor, Onit Alalouf, Ashwin Balakrishnan, Mike Heilemann, Ricardo Henriques, Yoav Shechtman Paper published in Small Methods, January 2025 Technologies: CARE () and ZeroCostDL4Mic () Funded by: Chan Zuckerberg Initiative (CZI), The Kavli Foundation, and The Wellcome Trust, CZI, EMBO, ERC, H2021 and H2022 DOI: 10.1002/smtd.202400672 |
|

|
DL4MicEverywhere - deep learning for microscopy made flexible, shareable and reproducible Iván Hidalgo-Cenalmor, Joanna W. Pylvänäinen, Mariana G. Ferreira, Craig T. Russell, Alon Saguy, Ignacio Arganda-Carreras, Yoav Shechtman, Guillaume Jacquemet, Ricardo Henriques, Estibaliz Gómez-de-Mariscal Paper published in Nature Methods, May 2024 Technologies: BioImage Model Zoo (), DL4MicEverywhere () and ZeroCostDL4Mic () Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1038/s41592-024-02295-6 |
|

|
The rise of data‐driven microscopy powered by machine learning Leonor Morgado, Estibaliz Gómez‐de‐Mariscal, Hannah S. Heil, Ricardo Henriques Review published in Journal of Microscopy, March 2024 Technologies: NanoJ () and NanoJ-Fluidics () Funded by: CZI, EMBO, ERC, FCT, H2021 and H2022 DOI: 10.1111/jmi.13282 |
|

|
Harnessing artificial intelligence to reduce phototoxicity in live imaging Estibaliz Gómez-de-Mariscal, Mario Del Rosario, Joanna W. Pylvänäinen, Guillaume Jacquemet, Ricardo Henriques Perspective published in Journal of Cell Science, February 2024 Technologies: BioImage Model Zoo (), CARE (), DeepBacs (), NanoJ-eSRRF (), NanoJ-SQUIRREL (), NanoJ-SRRF () and ZeroCostDL4Mic () Funded by: CZI, EMBO, ERC, H2021 and H2022 DOI: 10.1242/jcs.261545 |
|

|
This Microtubule Does Not Exist - Super‐Resolution Microscopy Image Generation by a Diffusion Model Alon Saguy, Tav Nahimov, Maia Lehrman, Estibaliz Gómez‐de‐Mariscal, Iván Hidalgo‐Cenalmor, Onit Alalouf, Ashwin Balakrishnan, Mike Heilemann, Ricardo Henriques, Yoav Shechtman Published in Small Methods, January 2024 Technologies: ZeroCostDL4Mic () Funded by: CZI, EMBO, H2021 and H2022 DOI: 10.1002/smtd.202400672 |
|

|
Nano-org, a functional resource for single-molecule localisation microscopy data Sandeep Shirgill, Daniel J Nieves, Jeremy A Pike, Mohammad A Ahmed, Mohammed HH Baragilly, Kylie Savoye, Jonathan Worboys, Khodor Hazime, Adrian Garcia, David J Williamson, Ricardo Henriques, Steven F Lee, Dylan M Owen Preprint published in bioRxiv, January 2024 Technologies: nano-org Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1101/2024.08.06.606779 |
|

|
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation Romain F. Laine, Hannah S. Heil, Simao Coelho, Jonathon Nixon-Abell, Angélique Jimenez, Theresa Wiesner, Damián Martínez, Tommaso Galgani, Louise Régnier, Aki Stubb, Gautier Follain, Samantha Webster, Jesse Goyette, Aurelien Dauphin, Audrey Salles, Siân Culley, Guillaume Jacquemet, Bassam Hajj, Christophe Leterrier, Ricardo Henriques Paper published in Nature Methods, November 2023 Technologies: CARE (), NanoJ (), NanoJ-eSRRF (), NanoJ-SQUIRREL (), NanoJ-SRRF () and Nuclear-Pores as references Funded by: CZI, EMBO, ERC, FCT, H2021, H2022, InnOValley and Wellcome Trust News: Photonics.com, The Science Times, Optics.org and Phys.org Blogs: Springer Nature Protocols and Methods Community DOI: 10.1038/s41592-023-02057-w |
|

|
Transertion and cell geometry organize the Escherichia coli nucleoid during rapid growth Christoph Spahn, Stuart Middlemiss, Estibaliz Gómez-de-Mariscal, Ricardo Henriques, Helge B. Bode, Séamus Holden, Mike Heilemann Preprint published in bioRxiv, October 2023 Technologies: CARE (), DeepAutoFocus (), DeepBacs () and ZeroCostDL4Mic () Funded by: EMBO, ERC, H2021 and H2022 DOI: 10.1101/2023.10.16.562172 |
|
News
- 2025-04-22: Blog BPoD - Biomedical Picture of the Day highlights Spahn et al. Nature Communications 2025 [external link]
- 2025-02-28: News outlet EUTOPIA highlights Saraiva et al. Nature Methods 2025 [external link]
- 2025-01-25: News outlet Tempo.pt highlights Saraiva et al. Nature Methods 2025 [external link]
- 2025-01-09: News outlet Nouvelles du monde highlights Saraiva et al. Nature Methods 2025 [external link]
- 2025-01-06: News outlet RTP highlights Saraiva et al. Nature Methods 2025 [external link]
- 2024-06-02: News outlet Labonline highlights Hidalgo-Cenalmor et al. Nature Methods 2024 [external link]
- 2024-05-28: News outlet MSN highlights Hidalgo-Cenalmor et al. Nature Methods 2024 [external link]
- 2024-05-28: News outlet AZoRobotics highlights Hidalgo-Cenalmor et al. Nature Methods 2024 [external link]
- 2024-05-27: News outlet Aamuset Kaupunkimedia highlights Hidalgo-Cenalmor et al. Nature Methods 2024 [external link]
- 2024-05-17: Blog news-medical.net highlights Hidalgo-Cenalmor et al. Nature Methods 2024 [external link]
- 2023-11-27: News outlet Photonics.com highlights Laine et al. Nature Methods 2023 [external link]
- 2023-11-20: News outlet The Science Times highlights Laine et al. Nature Methods 2023 [external link]
- 2023-11-16: News outlet Optics.org highlights Laine et al. Nature Methods 2023 [external link]
- 2023-11-14: Blog Springer Nature Protocols and Methods Community highlights Laine et al. Nature Methods 2023 [external link]
- 2023-11-14: News outlet Phys.org highlights Laine et al. Nature Methods 2023 [external link]